On scientific software - scope
Last time I discussed the idea that the notion of “quality scientific software” wasn’t very helpful, insofar as scientific code quality is a bit like beauty, being “in the eye of the beholder” - but I concluded with a few questions that IMHO are more important than “is it quality?”, being:
- Can I believe the results (produced by the software)?
- How were the results validated?, and
- Are they even useful?
However, before we get to answering these questions, there a few more layers of turtles to go through, in particular:
- What do we mean we talk about “the code” anyway?
- And why do the rest of you care?
To get a view on the second, I did a twitter poll about why we ask scientists to provide code when they write papers:
Broadly speaking most people think access to code matters because it aids in reproducibility, some people think seeing code makes “the method” more transparent, and a few people think that “good code” = “good science”. Only one person thinks that recording the code is just “good lab book practice” (making sure you know what was done by recording the parameters).
Actually, that’s not a fair characterisation of the poll, not least, because I had to ask “what was the most important”, rather than “prioritise the ones you think are important”. Also, here I’ve given context to what I meant that wasn’t there to those filling out the poll. However, it’s fairly indicative. In particular, I think it’s clear that most people think that the notion of code quality being indicative of good science is less important than other things, which include reproducibility and clarity of method. (There are also some really interesting comments in the poll answers, some of which I’ll come back to in a future post.)
But the poll was naughty in more than one way, not only did I make it a “pick one” (well twitter made me do it that way), but I also didn’t define what I meant by “the code” - which is fair, because in the main journal “requirements” don’t either. So, before we start going back up the turtle stack, let’s have a bash at defining what we mean by “the code”.
When many people think about “scientific code”, they have a mental model that is something like
- results = some function of (input data and code), where perhaps the function includes a bit of statistical nous.
I have a very different mental model:
- results = some function of (input data and code, and computing environment and possibly my digital laboratory) and code itself.
What do I mean by “my digital laboratory”? Well, there are our ways to get some input data for doing science:
- get data from someone else,
- make some measurements/observations, or
- do some simulations which generate numbers, or
- some combination of the other three.
All but the first involve code which generate numbers for analysis, so we need to think about not just the analysis code, but also the “digital laboratory code”.
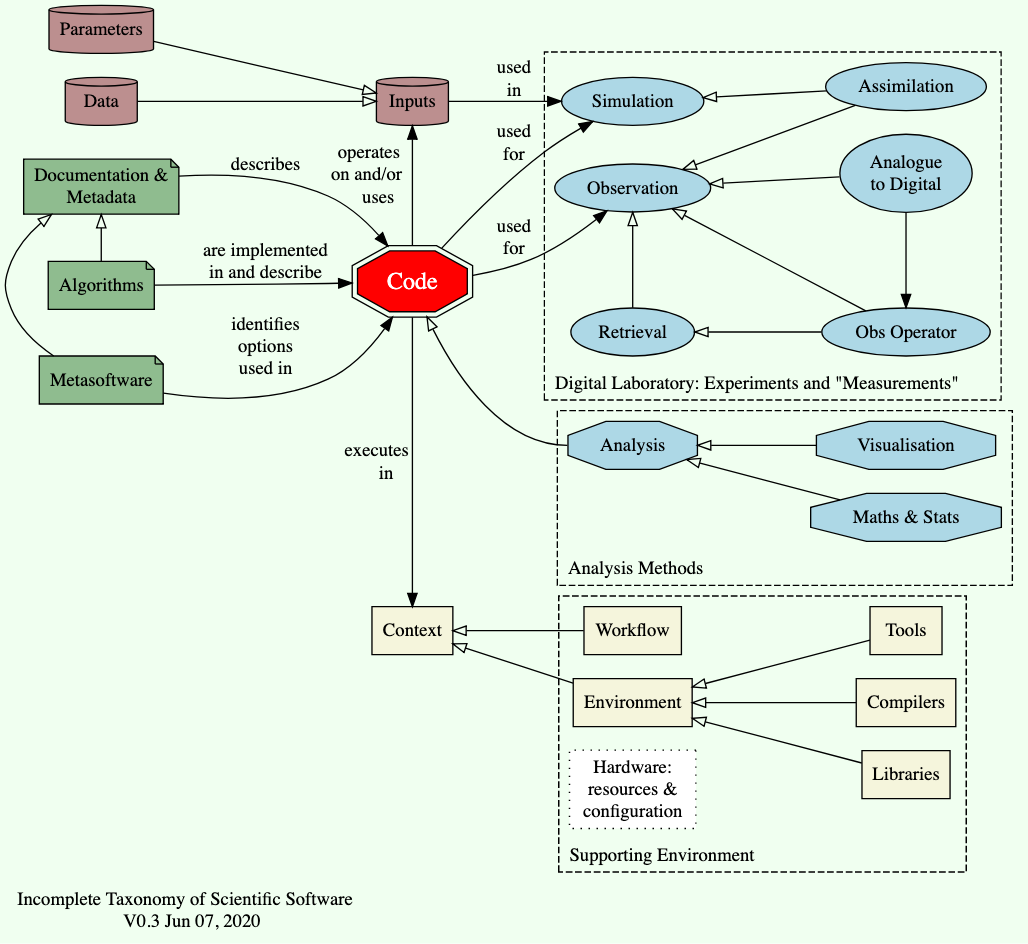
I’ve tried to put this all into one diagram:
At this point, hopefully you realise that we need to be a bit precise about what we mean by “code”, before we can talk about quality, or why we care about it.
In this diagram, the notation of an arrow with a hollow diamond means that the object at the base of the arrrow is a “specialisation” (loosely, a “type”) of the object at the head of the arrrow: i.e. analysis (code) is a type of code and visualisation (code) is a type of analysis code. At this point, hopefully you realise that we need to be a bit precise about what we mean by “code”, before we can talk about quality, or why we care about it.
You’ve probably also had enough for now, so we’re going to have to hold those three questions for a bit longer, and even longer before we get back to what I think about epidemiological modelling and its current use.
Comments on the story so far welcome on twitter.
(The figure was slightly corrected on the 7th of June, with no consequences to the story thus far.)